The database of Genotypes and Phenotypes (dbGaP) is a free resource that contains human data from a variety of large-scale studies. While you can’t view individual-level data without applying for controlled access, you can easily find dbGaP studies using the dbGaP Advanced Search (see screenshot below) and quickly filter studies based on study variables, molecular data type, study focus, NIH Institute, study consent, and more. Third-party annotations and mapping of phenotypic and study variables to controlled vocabularies allow you to search across studies. Once you find a study of interest, you can follow the Authorized Access link on records to apply for access.
Phenotypic and study variables include:
- Clinical measures (e.g., height, weight, blood pressure)
- Demographic information (e.g., age, gender, ethnicity)
- Sample information (e.g., analyte type, body site)
- Molecular data type (e.g., DNA sequence, genotypes, gene expression)
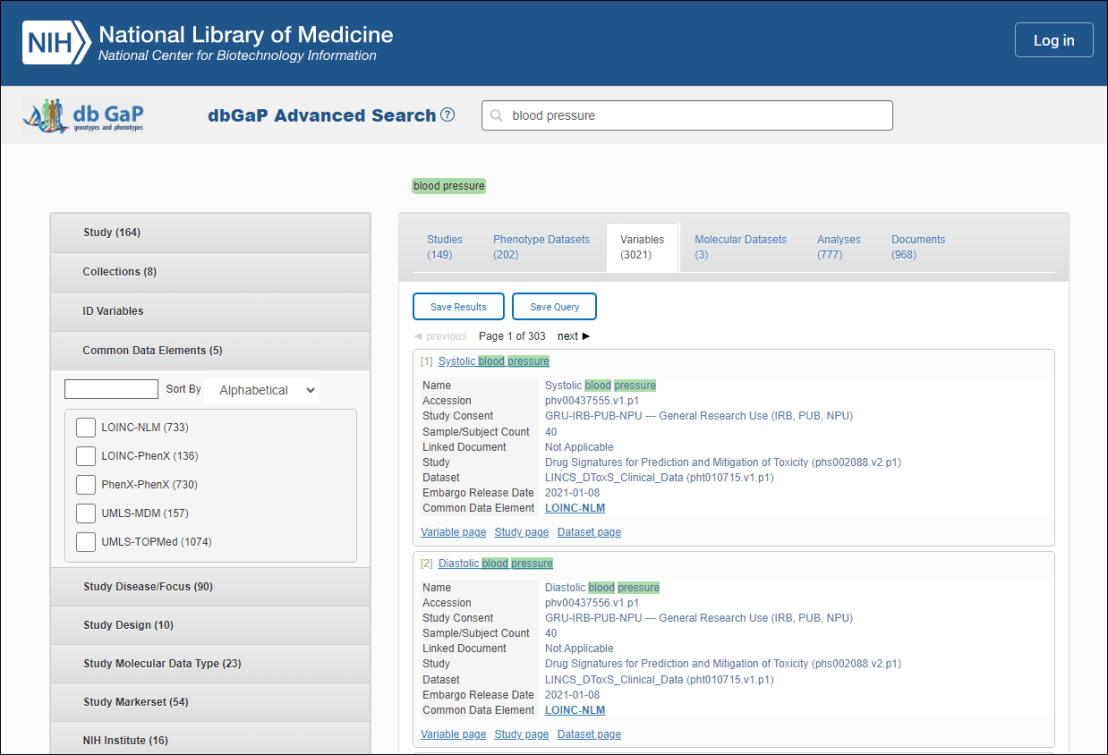
Outside groups have also provided additional annotations to help standardize and harmonize the terms used to describe phenotypes and variables. These include mapping to controlled vocabularies in the Unified Medical Language System UMLS Metathesaurus including the Logical Observation Identifiers Names and Codes (LOINC) vocabulary or directly to UMLS Concept Unique Identifiers (CUI).
Organizations doing the annotations include:
- NLM’s Lister Hill Center mapping to LOINC terms
- Medical Data Models mapping to UMLS CUI terms
- NHLBI’s TOPMed project mapping cardiovascular disease phenotypes to UMLS terms
- The PhenX project mapping to LOINC and PhenX terms
The annotated vocabularies will also be included in the dbGaP FHIR schema searchable through the dbGaP FHIR API.
Stay up to date
Follow us on Twitter @NCBI and join our mailing list to keep up to date with dbGaP and other NCBI news.
Questions?
If you have questions or would like to provide feedback, please reach out to us at info@ncbi.nlm.nih.gov.
